Module 1: Actin Filaments
G-actin assembles head-to-tail into polar, double-helical F-actin filaments that drive cell shape, motility, contraction, and many membrane-remodeling events. We build the Oosawa-Asakura kinetic theory, derive the critical concentration and treadmilling, then meet the regulators: Arp2/3 (branching), formins (straight elongation), cofilin/ADF (severing), CapZ (capping), and the NPFs (WASP/WAVE) that integrate the network with Rho-GTPase signaling.
1. G-actin, F-actin, and the ATP Switch
Actin is a 375-residue, 42 kDa ATPase whose monomer (G-actin) contains a deep cleft that binds either ATP or ADP and a divalent cation (Mg2+ in cells, Ca2+ in typical biochemistry). Kabsch et al. (1990) solved the atomic structure of G-actin·DNase I; subsequent structures by Otterbein et al. (2001) and Dominguez (2004) revealed that ATP binding clamps the cleft closed, while ADP binding allows it to open —a conformational switch that tunes monomer-monomer interfaces.
When G-ATP-actin concentration exceeds the critical concentration, monomers polymerize into a right-handed, double-helical filament (F-actin) of diameter 7 nm with a crossover repeat of 36 nm. Each subunit hydrolyzes its bound ATP to ADP-Pi within a few seconds of incorporation; Pi release follows with a half-life of \(\sim 5\)minutes, giving the filament a “nucleotide clock” that locally marks subunit age.
\[\text{G-ATP} \;\xrightarrow{k_{\text{on}}}\; \text{F-ATP} \;\xrightarrow{k_{\text{hyd}}}\; \text{F-ADP-P}_i \;\xrightarrow{k_{\text{P}_i}}\; \text{F-ADP}\]
Typical rates: \(k_{\text{hyd}} \approx 0.3\,\text{s}^{-1}\),\(k_{\text{P}_i} \approx 3\times 10^{-3}\,\text{s}^{-1}\)(Carlier & Pantaloni 1986).
The three nucleotide states have distinct affinities at each filament end. This is the single most important fact of actin biochemistry: it generates the polar asymmetry of the filament and drives treadmilling.
2. Oosawa-Asakura Kinetics
Oosawa & Asakura (1975) modeled actin polymerization as a two-step process: (a) a slow, unfavorable nucleation of a small oligomer (dimer, trimer), and (b) a fast elongation by end-wise monomer addition. Writing \([M]\) for the monomer concentration and \([E]\) for the density of growing ends,
\[\frac{dF}{dt} = (k_{\text{on}}[M] - k_{\text{off}})\,[E], \qquad \frac{d[E]}{dt} = k_{\text{nuc}}\,[M]^{n^*}\]
\(F\) = polymer mass (monomers incorporated), \(n^* \approx 3\) = nucleus size,\(k_{\text{nuc}}\) = effective pre-equilibrium constant.
Critical concentration
At steady state \(dF/dt = 0\) and \([M] = C_c = k_{\text{off}}/k_{\text{on}}\). For actin at the barbed end with ATP-G, \(C_c^+ \approx 0.12\,\mu\)M; at the pointed end with ADP-G, \(C_c^- \approx 0.6\,\mu\)M (Pollard 1986). Because these are unequal, the polymer has a net polarity: at any bulk monomer concentration\(C_c^+ < [M] < C_c^-\), the plus end grows while the minus end shrinks.
Nucleation lag
Because nucleation scales as \([M]^{n^*}\), the onset of polymerization shows a marked concentration-dependent lag. The characteristic time is\(\tau_{\text{lag}} \sim 1/\sqrt{k_{\text{on}}\,k_{\text{nuc}}\,[M]^{n^*+1}}\)(Oosawa & Kasai 1962). This lag is abolished in vivo by cellular nucleators—Arp2/3, formins, tandem-monomer binders—which template the nucleus directly.
Polymer mass vs. total actin: the hockey stick
3. Treadmilling: Polar Asymmetry and ATP Hydrolysis
The polarity of F-actin is commonly visualized by decoration with myosin-S1 heads, which bind at a fixed azimuthal angle producing a barbed (+) end (wide) and a pointed (−) end (narrow). The two ends have different rate constants, especially for the different nucleotide states:
| end / state | \(k_{\text{on}}\) (uM\(^{-1}\)s\(^{-1}\)) | \(k_{\text{off}}\) (s\(^{-1}\)) | \(C_c\) (uM) |
|---|---|---|---|
| Barbed (+), ATP | 11.6 | 1.4 | 0.12 |
| Barbed (+), ADP | 3.8 | 7.2 | 1.9 |
| Pointed (−), ATP | 1.3 | 0.8 | 0.6 |
| Pointed (−), ADP | 0.16 | 0.27 | 1.7 |
Treadmilling is the steady state in which the polymer has a bulk monomer concentration between the two ends’ critical concentrations, so that net assembly occurs at the barbed end and net disassembly at the pointed end, with subunits flowing through the filament along their lifetime. ATP hydrolysis drives this non-equilibrium steady state (it costs \(\sim 1\) ATP per subunit of treadmilled flux).
\[v_{\text{tread}} = k_{\text{on}}^+\,[M]_{\text{ss}} - k_{\text{off}}^+ = -\bigl(k_{\text{on}}^-\,[M]_{\text{ss}} - k_{\text{off}}^-\bigr)\]
The shared steady-state monomer concentration is \([M]_{\text{ss}} \approx 0.1\text{--}0.3\,\mu\)M in resting cells.
Treadmilling: subunit flux through a polar filament
4. Arp2/3 Complex & Dendritic Nucleation
The Arp2/3 complex (seven subunits: Arp2, Arp3, and ARPC1-5) is the universal branched-actin nucleator. It is activated by Nucleation-Promoting Factors (NPFs: WASP, N-WASP, WAVE/Scar, WHAMM, JMY), which deliver a G-ATP-actin monomer to a mother filament and induce a conformational change that assembles a stable branch point. Branches form at a characteristic angle of \(70 \pm 7^\circ\) (Mullins, Heuser & Pollard 1998; Svitkina & Borisy 1999).
Dendritic network model
Pollard & Borisy’s dendritic-nucleation/array-treadmilling model (2003) and Blanchoin, Sykes, Plastino & Boujemaa-Paterski’s review (2014) organize the lamellipodium into a cycle:
- PIP2 and Rac-GTP activate WASP/WAVE NPFs near the membrane.
- Active NPF recruits Arp2/3 and G-ATP-actin to the side of an existing mother filament.
- A new daughter filament nucleates at 70\(^\circ\) to the mother, barbed end outward.
- Barbed-end elongation (6–14 \(\mu\)m/min) pushes the membrane forward.
- Capping protein (CapZ) caps filaments after a few \(\mu\)m, preventing bouquet collapse.
- ATP hydrolyzes; cofilin preferentially severs ADP-F-actin at the rear, recycling monomers.
\[\langle L_{\text{filament}} \rangle = \frac{v_{\text{elong}}}{k_{\text{cap}}}\]
With \(v_{\text{elong}} \approx 0.3\,\mu\)m/s and \(k_{\text{cap}} \approx 3\,\text{s}^{-1}\),\(\langle L \rangle \approx 0.1\,\mu\)m—consistent with EM measurements.
Arp2/3 dendritic branched network (schematic)
5. Formins: Processive Straight Elongation
Formins (mDia1/2/3, Dia, FMNL, DAAM, INF2, FHOD, Cappuccino) nucleate unbranched actin filaments. They contain an FH1 domain rich in polyproline tracks that captures profilin-actin, and an FH2 donut dimer that encircles the barbed end and remains processively attached as the filament elongates.
\[v_{\text{formin}} = k_{\text{on}}^{\text{PA}} [\text{PA}] - k_{\text{off}}\]
The formin uses an FH1-tethered profilin-actin (PA) pool. Effective \(k_{\text{on}}^{\text{PA}}\)can exceed the diffusion-limited rate by an order of magnitude.
Because formins processively track the barbed end, a single formin adds thousands of subunits without dissociating (Kovar & Pollard 2004; Michelot & Drubin 2011). Formin-elongated filaments are linear, not branched, and populate stress fibers, filopodia, and the contractile ring.
6. Cofilin/ADF: Severing Old Filaments
Cofilin (ADF/cofilin family) preferentially binds F-ADP-actin and changes the filament twist by \(\sim 5^\circ\) per subunit, weakening longitudinal contacts. At high occupancy, cofilin severs filaments at the cofilin/bare-actin boundary (McGough 1997; De La Cruz 2005); at saturating levels it debranches Arp2/3 junctions (Chan, Beltzner & Pollard 2009).
Because cofilin prefers aged (ADP) filaments, the lamellipodial actin network is severed and recycled specifically at the rear of the network, which is the oldest. This sets up the dendritic treadmill: assembly at the front, severing and monomer recycling at the rear.
\[k_{\text{sev}} \;=\; k_0 \,\theta_{\text{cof}}(1-\theta_{\text{cof}})\]
Severing rate peaks at intermediate cofilin occupancy \(\theta_{\text{cof}} \approx 0.5\)—the boundary-density dependence.
7. Capping Proteins & Nucleation-Promoting Factors
CapZ (alpha/beta heterodimer, also called capping protein CP) binds F-actin barbed ends with nanomolar affinity and blocks both addition and loss. In lamellipodia, CapZ is the primary regulator of filament length (\(\approx 0.1\) \(\mu\)m). Tropomodulin caps the pointed end in sarcomeres and erythrocyte membrane skeletons.
Nucleation-Promoting Factors couple upstream signals to Arp2/3 activation. The common architecture is an autoinhibited protein that opens only upon binding of activated Rho-family GTPases, PIP2, and SH3 adaptors. The C-terminal VCA module then recruits Arp2/3 and G-actin.
- WASP / N-WASP: activated by Cdc42-GTP & PIP2; filopodia and endocytic patches.
- WAVE/Scar complex: activated by Rac-GTP; lamellipodia.
- WHAMM: ER tubulation and autophagy.
- JMY: nuclear and cytoplasmic, p53 coactivator.
8. Cellular Architectures Built from Actin
Lamellipodium
Flat, protrusive actin veil at the leading edge; Arp2/3 branched network; Rac/WAVE driven. Advance rate 0.1–1 \(\mu\)m/s.
Filopodium
Finger-like parallel bundle of 10–30 filaments; fascin bundler, formin-nucleated; Cdc42/WASP driven. Explores substrate and matrix.
Stress fiber
Antiparallel actin/myosin bundle anchored at focal adhesions; contractile, mechanosensitive. Rho/mDia driven, Ser-19 MLC phosphorylation.
Contractile ring
Equatorial actomyosin bundle driving cytokinetic furrow; anillin, septins, formin (mDia2). Constriction speed 0.3 \(\mu\)m/min (human cells).
Cell cortex
Thin (200–500 nm) isotropic actomyosin mesh beneath the membrane; sets cell surface tension \(\sim 10^{-3}\) N/m.
Muscle thin filament
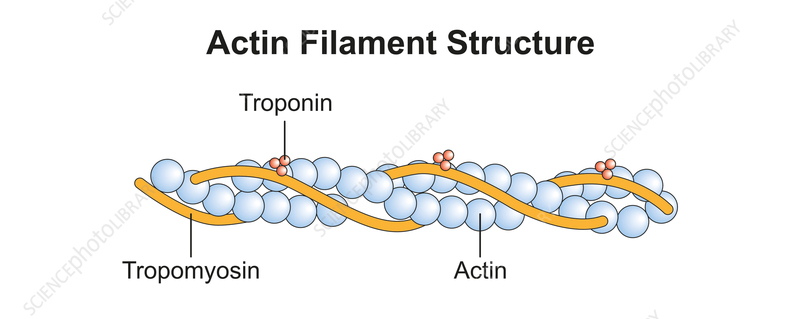
Ordered F-actin decorated with tropomyosin + troponin complex; Ca-regulated myosin binding; sarcomere length 2.0–2.5 \(\mu\)m.
9. Free-Energy Accounting of Actin Polymerization
The free energy of monomer addition to a filament end is \(\Delta G = k_B T \ln(C_c / [M])\). At physiological \([M]_{\text{ss}} \approx 0.2\,\mu\)M and\(C_c^+ \approx 0.12\,\mu\)M, addition is slightly exergonic (~0.5 \(k_B T\)), while pointed-end dissociation is similarly exergonic. Hydrolysis of ATP to ADP + Pion the filament releases about 13 \(k_B T\), much of which is dissipated; the remainder sets up the free-energy difference that sustains treadmilling.
\[\Delta G_{\text{tread}} \;=\; k_B T \ln\frac{C_c^-}{C_c^+} \;\approx\; k_B T \ln\frac{0.6}{0.12} \;\approx\; 1.6\,k_B T\,\text{per subunit}\]
Treadmilling dissipates about one ATP (\(\sim 20\,k_B T\)) per subunit cycled, most of it as heat. Only a small fraction is captured as directed mechanical work (e.g. against membrane tension).
Force generation by polymerization
Peskin, Odell & Oster’s (1993) Brownian ratchet model explains how an assembling filament pushes on a fluctuating load (e.g. the leading-edge membrane). When the fluctuating load transiently moves away by \(\delta\), a monomer fits in, and the step is rectified. The stall force is
\[F_{\text{stall}} \;=\; \frac{k_B T}{\delta}\,\ln\frac{[M]}{C_c}\]
With \(\delta = 2.7\) nm and \([M]/C_c \sim 50\),\(F_{\text{stall}} \approx 6\) pN per filament — enough for the hundreds-of-pN load of a leading edge distributed across \(\sim 100\) filaments/\(\mu\)m.
Profilin and sequestration
In cells, most G-actin is bound to profilin, a small 14 kDa protein that occludes the pointed face of G-actin. The profilin-actin complex can add to the barbed end but not the pointed end, biasing polymerization strongly in the plus direction. Profilin also accelerates nucleotide exchange on G-actin and presents G-actin to formin FH1 domains. Thymosin-β4, by contrast, simply sequesters G-actin, providing a buffer pool.
Simulation 1: Two-Ended Polymerization & Treadmilling
Explicit integration of a four-reaction scheme (two ends \(\times\) two nucleotide states) with ATP hydrolysis, Pi release, and exchange on free monomer. The system approaches a treadmilling steady state in which barbed-end influx equals pointed-end outflux, sustained by ATP hydrolysis.
Click Run to execute the Python code
Code will be executed with Python 3 on the server
Simulation 2: Arp2/3 Dendritic Branching Network
Stochastic 2-D simulation of an expanding lamellipodial network: filaments elongate, stochastically produce 70\(^\circ\) Arp2/3 branches, and are terminated by capping with rate constant\(k_{\text{cap}}\). The simulation reproduces the observed mean-length\(v_{\text{elong}}/k_{\text{cap}}\) scaling and the dendritic geometry of real lamellipodia (Svitkina & Borisy 1999).
Click Run to execute the Python code
Code will be executed with Python 3 on the server
Key References
• Oosawa, F. & Asakura, S. (1975). Thermodynamics of the Polymerization of Protein. Academic Press.
• Pollard, T.D. (1986). “Rate constants for the reactions of ATP- and ADP-actin with the ends of actin filaments.” J. Cell Biol., 103, 2747–2754.
• Pollard, T.D. & Borisy, G.G. (2003). “Cellular motility driven by assembly and disassembly of actin filaments.” Cell, 112, 453–465.
• Blanchoin, L., Boujemaa-Paterski, R., Sykes, C., & Plastino, J. (2014). “Actin dynamics, architecture, and mechanics in cell motility.” Physiol. Rev., 94, 235–263.
• Mullins, R.D., Heuser, J.A., & Pollard, T.D. (1998). “The interaction of Arp2/3 complex with actin.” PNAS, 95, 6181–6186.
• Svitkina, T.M. & Borisy, G.G. (1999). “Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia.” J. Cell Biol., 145, 1009–1026.
• Kovar, D.R. & Pollard, T.D. (2004). “Insertional assembly of actin filament barbed ends in association with formins produces piconewton forces.” PNAS, 101, 14725–14730.
• Carlier, M.-F. & Pantaloni, D. (1986). “Direct evidence for ADP-Pi-F-actin as the major intermediate in ATP-actin polymerization.” Biochemistry, 25, 7789–7792.
• Spudich, J.A. & Watt, S. (1971). “The regulation of rabbit skeletal muscle contraction.” J. Biol. Chem., 246, 4866–4871.
• Kabsch, W. et al. (1990). “Atomic structure of the actin:DNase I complex.” Nature, 347, 37–44.
• Kuhn, J.R. & Pollard, T.D. (2005). “Real-time measurements of actin filament polymerization by TIRF microscopy.” Biophys. J., 88, 1387–1402.
• Chan, C., Beltzner, C.C., & Pollard, T.D. (2009). “Cofilin dissociates Arp2/3 complex and branches from actin filaments.” Curr. Biol., 19, 537–545.
• De La Cruz, E.M. (2005). “Cofilin binding to muscle and non-muscle actin filaments.” J. Mol. Biol., 346, 557–564.
• Michelot, A. & Drubin, D.G. (2011). “Building distinct actin filament networks in a common cytoplasm.” Curr. Biol., 21, R560–R569.
• Alberts, B. et al. (2015). Molecular Biology of the Cell, 6th ed., Garland Science.